SNPs in the promoter region may affect the expression of a gene either by creating or removing regulatory elements. SNPs in the 3-untranslated region of a transcript may influence the stability and give higher or lower levels of expression based on the genotype. We are working on identifying such SNPs in genes that show high interindividual variation in expression in microarray experiments but little variation in expression in the same individual. The genes selected are all part of different breast tumour subclasses and the SNPs that affect gene expression could be used as diagnostic and prognostic markers. Data mining for SNPs is performed using the web based databases Ensembl (
http://www.ensembl.org/), TSC (The SNP Consortium) (
http://snp.cshl.org/) and the Genome Browser (
http://genome.ucsc.edu/). Position, rs number (unique identification number) and functionality (coding, non-coding, intronic, 3'or 5'UTR) of each SNP are thus obtained.

| 
|
A.
| B.
|
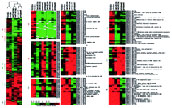
Figure. Genetic and environmental factors account for the difference in homogeneity of clustering of gene expression between outbred humans (A.) and inbred rats (B.) Identification of the genetic factors is in the focus of this project.
Key member: Silje Nordgard


