A basic study of the cell cycle by mass cytometry, with implications for exclusion of non-single cell events as ion cloud fusions and aggregates.

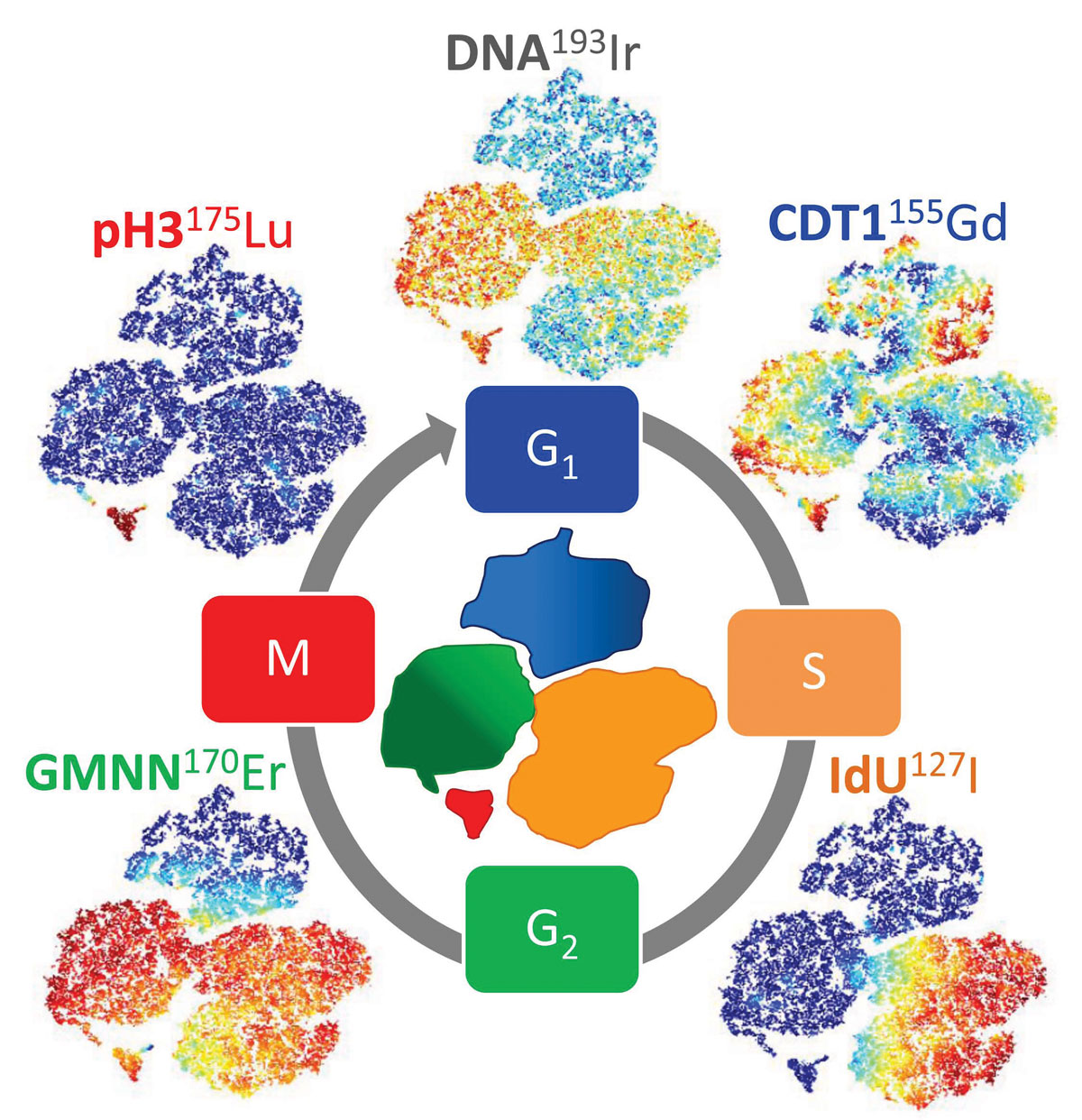
A recent study from the Sections of Core Facilities/Radiation Biology/Tumor Immunology shows that the cell cycle can be completely resolved by mass cytometry, including subdivision of the G1 and G2 phases. Only a few markers are needed, and the remaining >50 parameters are free to measure the expression of other (phospho)proteins of interest. Stratification of cells into different cell cycle phases also made exclusion of non-single cell events possible, solving another problem in mass cytometry.
The cell cycle has been intensively studied for the last half century, particularly in cancer research. No tumors can arise without uncontrolled cell proliferation, and many oncogene- and tumor suppressor gene products have impact on cell cycle progression.
Cell cycle studies have mostly been performed by methods giving single cell resolution, e.g. flow cytometry. Mass cytometry is a relatively new method that allows for assessment of up to 60 parameters, i.e. many more than achievable by flow cytometry, thus approaching high content screening on the single cell level. However, methods for studying the cell cycle by mass cytometry cannot easily be translated from flow cytometry protocols, there is e.g. no good probe for measuring DNA content accurately by mass cytometry.
Links:
The article:
Cell Cycle Analysis and Relevance for Single-Cell Gating in Mass Cytometry.
Rein ID, Notø HØ, Bostad M, Huse K, Stokke T.
Cytometry A. 2020 Jan 14. doi: 10.1002/cyto.a.23960.
PMID: 31943748
Flow Cytometry Core Facilities at Oslo University Hospital
Trond Stokke's project group - Molecular Radiation Biology group
