Welcome to the home page of Jorrit Enserink's group Cancer Molecular medicine
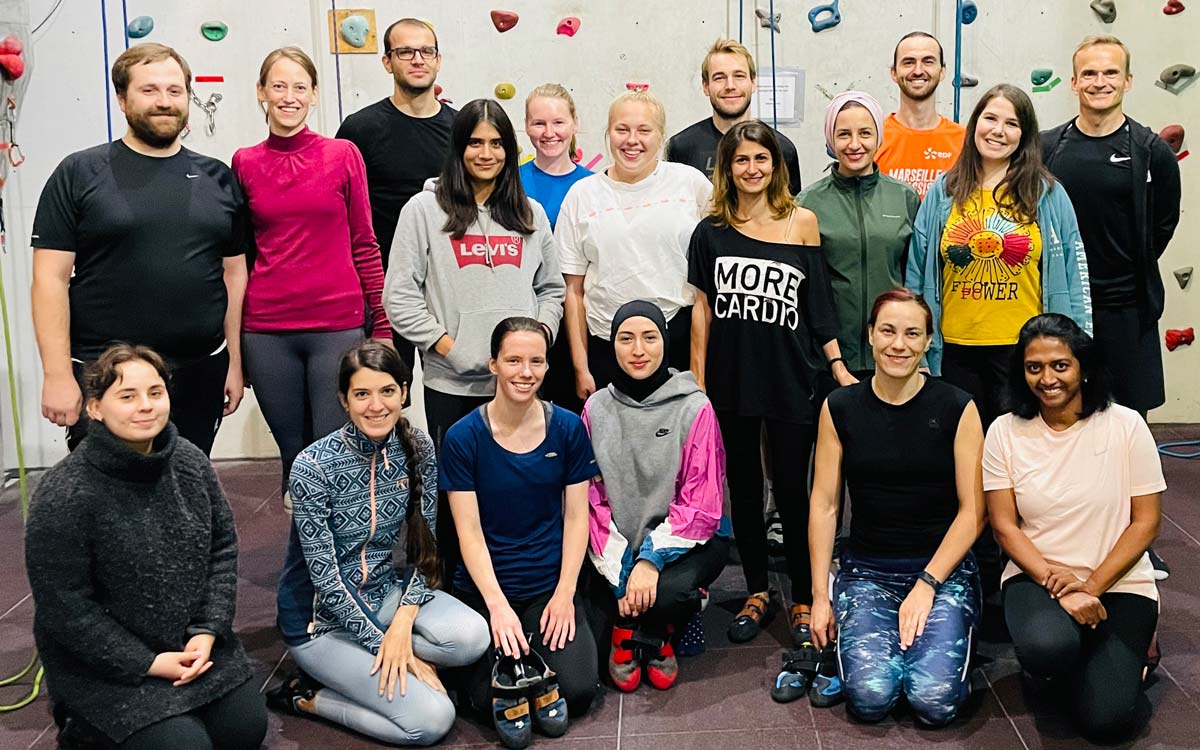
The Enserink group is a research group with people from eleven different countries, and currently consists of the group leader, two senior researcher, seven post-docs, three PhD students and a varying number of undergraduate students. We make use of model organisms as well as primary material from cancer patients to study several basic problems in cancer.

Click on the images below for more information on our research projects:
Personalized medicine and oncogene addiction
We are funded by the following organizations:
 |
 |